Heatmap graphpad prism6/24/2023  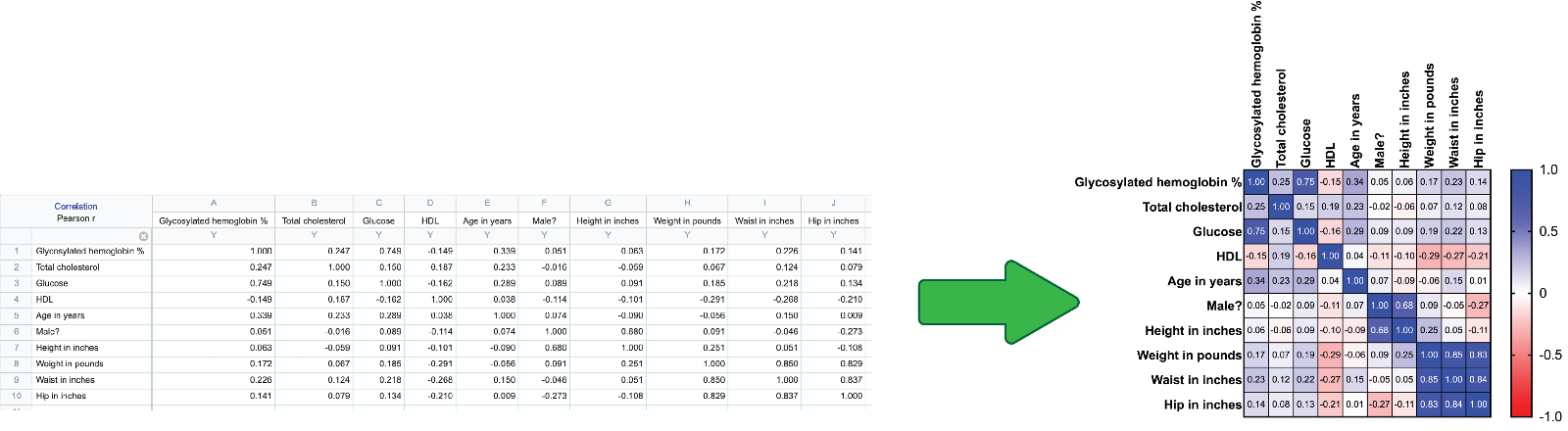
Gene ID|baseMean|Base Mean Sample 1|Base Mean Sample 2| Normalized value|log2 fold-change | p value | q value | So something like this should be fine for both DESeq and EdgeR tables: Thank you dpryan for all the great advice. If someone could kindly help me on how I could go about this. In total I hope to have 3 Heatmaps with the specific genes of interests only: The p values and log information seem to indicate high values for the genes that I am interested in. I have 2 replicates for Organ A and 2 replicates for Organ B. I have already used Tuxedo pipeline to create a heat map using CummeRbund for the species. Or is there any better method to analyze these genes via EdgeR and DeSeq. Now I was wondering how could I possibly create heat map based on the genes that I interested based on EdgeR and DeSeq. Edited/removed no_feature, ambiguous, too_low_aQual:, not_aligned, alignment_not_unique in the output file and added Organs headers for each rep.ĥ) Used EdgeR and DeSeq respectively on Iplant Collaborative. (offline)Ĥ) Used Iplant Collaborative Join tab delimited to merge the Counts into one file. 
I have done the following:ġ) Tophat2 (provided Ensembl GTF and Fasta files)Ģ) Used Samtools to convert the files to Sam format (offline)ģ) HTseq and used the same GTF file from Tophat2. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
